What is the difference between Maxam Gilbert and Sanger method?
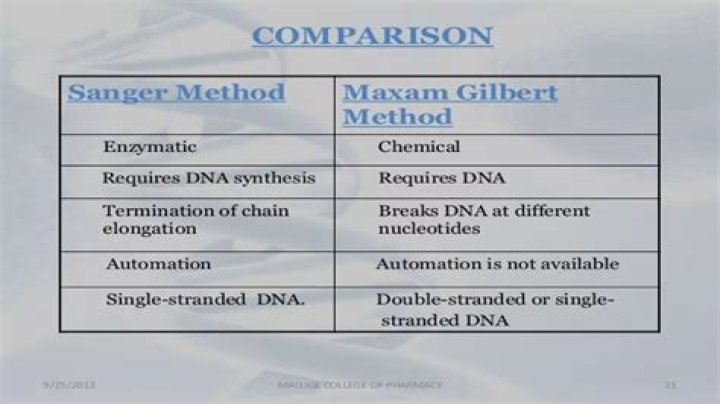
Maxam–Gilbert technique depends on the relative chemical liability of different nucleotide bonds, whereas the Sanger method interrupts elongation of DNA sequences by incorporating dideoxynucleotides into the sequences. The chain termination method is the method more usually used because of its speed and simplicity.
What is better than Sanger sequencing?
next-generation sequencing (NGS) technologies are similar. While the Sanger method only sequences a single DNA fragment at a time, NGS is massively parallel, sequencing millions of fragments simultaneously per run. This high-throughput process translates into sequencing hundreds to thousands of genes at one time.
What is Maxam Gilbert sequencing used for?
Maxam–Gilbert sequencing was the first widely adopted method for DNA sequencing, and, along with the Sanger dideoxy method, represents the first generation of DNA sequencing methods.
Why was the early Maxam Gilbert method of DNA sequencing not a favorite among early DNA scientists?
First, it was time consuming. And that was supposing that everything went well on the first try. A lot of steps in the method could cause problems: the radioactive labeling process, the cleavage reactions, the gel set up, the electrophoresis, and the X-ray film developer.
Is PCR used in Maxam Gilbert method?
Direct sequencing of polymerase chain reaction (PCR) products by using the Maxam-Gilbert method is described. In this method, one of the primers is end labeled. Thus it is possible to sequence the reaction product directly following purification using this chemical method.
Which type of DNA cleavage is done in the Maxam and Gilbert method?
4. Which type of DNA cleavage is done in the Maxam Gilbert method? Explanation: The Maxam Gilbert method of sequencing was devised in 1977. It uses a variety of chemical reagents to bring about base-specific cleavage of DNA.
How is Illumina better than Sanger?
The primary practical difference between Sanger sequencing and next generation sequencing is the yield of sequence data. Illumina’s sequencing machine can produce up to 20 mega bases (Mb) per hour with a read length of 100 bases from both ends of the template.
Which type of cleavage is done in Maxam Gilbert method?
When was Maxam Gilbert sequencing invented?
1977
The Maxam–Gilbert method was produced in 1975/76 and the paper detailing it was published in 1977. Gilbert’s students reached a point where they managed to sequence a 5000 base-long plasmid.
Which type of cleavage is done in Maxam-Gilbert method?
How many types of chemical treatments are required in Maxam-Gilbert method of sequencing?
There are four chemical cleavage reactions at the core of the Maxam and Gilbert sequencing system. The figure below left shows an example from these reactions, the reaction cleaving specifically at guanine. The other three reactions cleave at G+A, C+T, or C.
What is the difference between Maxam-Gilbert and Sanger sequencing?
The Maxam-Gilbert method is based on chemical modification of DNA and subsequent cleavageof the DNA backbone at sites adjacent to the modified nucleotides. Sanger sequencing uses specific chain-terminating nucleotides (dideoxy nucleotides) that lack a 3′-OHgroup.
What is the difference between Sanger dideoxy and Maxam Gilbert?
The Chain Termination Method (also known as the Sanger dideoxy method after its inventor). Maxam–Gilbert technique depends on the relative chemical liability of different nucleotide bonds, whereas the Sanger method interrupts elongation of DNA sequences by incorporating dideoxynucleotides into the sequences.
Who developed the Sanger-Gilbert method?
In the 1970s Frederick Sanger , and Walter Gilbert along with Allan Maxam developed the Sanger Sequencing and Maxam-Gilbert Sequencing methods respectively. Frederick Sanger won 2 Nobel prizes in chemistry. One in 1958 for his work in structure of proteins especially insulin.
What chemicals are used in Maxam Gilbert sequencing?
Maxam gilbert method uses base specific chemicals to break DNA at specific bases. Two common chemicals named dimethyl sulphate and hydrazine chemicals are used to selectively attack purines and pyrimidines, respectively. Maxam Gilbert sequencing method is performed via several steps as follows.

